CUDA Spotlight: Dylan Jackson By Calisa Cole, posted May 24, 2013 GPU-Accelerated Model Fitting of Interferometric Imagery for Label-Free BiosensingThis week's Spotlight is on Dylan Jackson of MIT Lincoln Laboratory. Previous to joining MIT-LL, Dylan was a student at Boston University working in the area of optical bionsensors. His master's degree project (Fall 2012) explored the application of GPU computing to the LED-based Interferometric Reflectance Imaging Sensor (IRIS) system. This interview is part of the CUDA Spotlight Series. Q & A with Dylan JacksonNVIDIA: Dylan, tell us about your master's degree project ("Improved Performance of CaFE and IRIS Model Fitting Using CUDA").
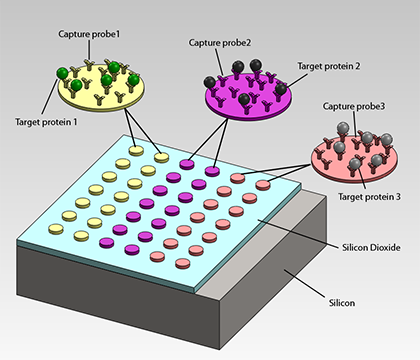
Optical biosensors are known to be accurate and reliable tools for measuring and monitoring the binding and unbinding of target proteins. Recently, new techniques and technologies have emerged to enable high-throughput biosensing with smaller, lower cost, and less complex systems. In particular, the LED-based Interferometric Reflectance Imaging Sensor (IRIS) has been demonstrated as a viable alternative to previously established high-end biosensors. The IRIS technique utilizes optical interference on a silicon dioxide on silicon slide to directly image and measure the protein density on the surface with a high degree of precision.
While IRIS enables high-throughput detection, new processing challenges were introduced as a result of measures to reduce cost and increase robustness. In particular, the use of LEDs instead of a tunable laser greatly increases the computational complexity of the reflectance model. We are using the Levenberg-Marquardt algorithm for fitting the data to a reflectance model. Since this is an iterative algorithm, increasing the complexity of the model compounds the processing power and time required for convergence. The pixel-based nature of the data lent itself well to data level parallelism, so it was natural to explore single instruction, multiple data (SIMD) architectures to improve processing performance. Since CUDA enables parallel computing on GPUs it seemed to be an ideal platform for exploiting data level parallelism on a single computer. For this project, I developed a CUDA implementation of the IRIS model, yielding significant performance improvements over the current MATLAB implementation. NVIDIA: Talk about key challenges in the field of biosensors. On the technical side, many current technologies can detect how much molecular binding occurs on the microarray, but this comes without knowledge of how much probe material initially existed on any given node. For example, one spot could show very little binding but there would be no way to tell how many probe molecules were there to begin with. Was there 100% binding to very few probes or 0.01% binding to a great number of probes? IRIS can be used to measure the amount of probe material to yield more accurate results. Switching IRIS from using a tunable laser to LEDs was an important step in the direction of reducing cost and improving reliability, but the increase in model complexity significantly increased the computational requirements. A computing grid took 15 to 20 minutes to iteratively fit a single multi-spectral image stack to the reflectance model. A single computer would need a whole day to do the same processing. If you want to perform rapid diagnosis in a remote area, you may only have access to one or two computers. In that situation you can’t afford to tie up a computer for an entire day just to process a single sensor. NVIDIA: What made you decide to look at CUDA? I reasoned that a GPU would be an economical way to solve the processing time and resource problem by acting as a coprocessor to the CPU. This would free up local CPU resources for other operations while taking advantage of parallel computing hardware, and better yet, it would only require a single computer. NVIDIA: In what specific ways did you leverage CUDA? One potential problem with applying CUDA to iteratively fitting multiple point sets is that each pixel could converge to a fit on a different iteration. If a thread is assigned to each pixel – an intuitive execution configuration for image processing – resources can be wasted processing completed pixels due to warp granularity. I found that assigning a block to each pixel allowed me to efficiently perform the necessary calculations for fitting, while quickly skipping over pixels that had already been flagged as converged. NVIDIA: What are some potential real-world applications of LED-IRIS? NVIDIA: What are you working on at MIT Lincoln Laboratory? NVIDIA: How did you become interested in this field? NVIDIA: Tell us about your band, Remedy Transmission. Bio for Dylan JacksonDylan currently works full time at MIT Lincoln Laboratory and plays in an original progressive rock band in his free time. His M.S. concentration was in electromagnetics and photonics. At MIT-LL he began developing low power, wireless sensor nodes. His work has since shifted to the development of advanced optical (MWIR, LWIR) systems and CUDA programming for real-time image processing. He holds B.S. and M.S. degrees from Boston University. Relevant Links Contact Info |